
SEGWE calculator: Diffusion coefficient and molecular weight estimation | Manchester NMR Methodology Group
Hydrodynamic Radii of Intrinsically Disordered Proteins Determined from Experimental Polyproline II Propensities | PLOS Computational Biology
Measurements of hydrodynamic radius and effective electrical charge of... | Download Scientific Diagram

SAXSMoW 2.0: Online calculator of the molecular weight of proteins in dilute solution from experimental SAXS data measured on a relative scale - Piiadov - 2019 - Protein Science - Wiley Online Library

Calculation of Hydrodynamic Properties of Globular Proteins from Their Atomic-Level Structure: Biophysical Journal

Monitoring Protein Global and Local Parameters in Unfolding and Binding Studies: The Extended Applicability of the Diffusion Coefficient─Molecular Size Empirical Relations | Analytical Chemistry
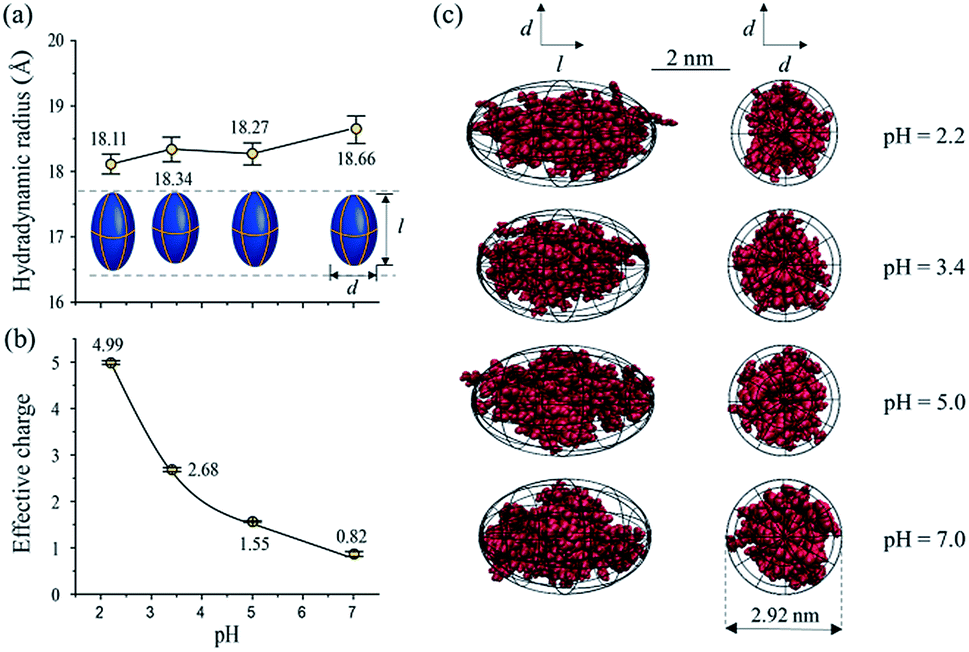
Structure and effective charge characterization of proteins by a mobility capillary electrophoresis based method - Chemical Science (RSC Publishing) DOI:10.1039/C9SC02039J
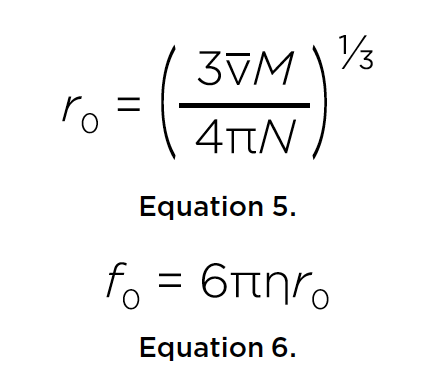
Temperature dependence of hydrodynamic radius of an intrinsically disordered protein measured in the Optima AUC analytical ultracentrifuge.
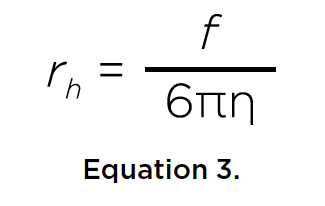
Temperature dependence of hydrodynamic radius of an intrinsically disordered protein measured in the Optima AUC analytical ultracentrifuge.
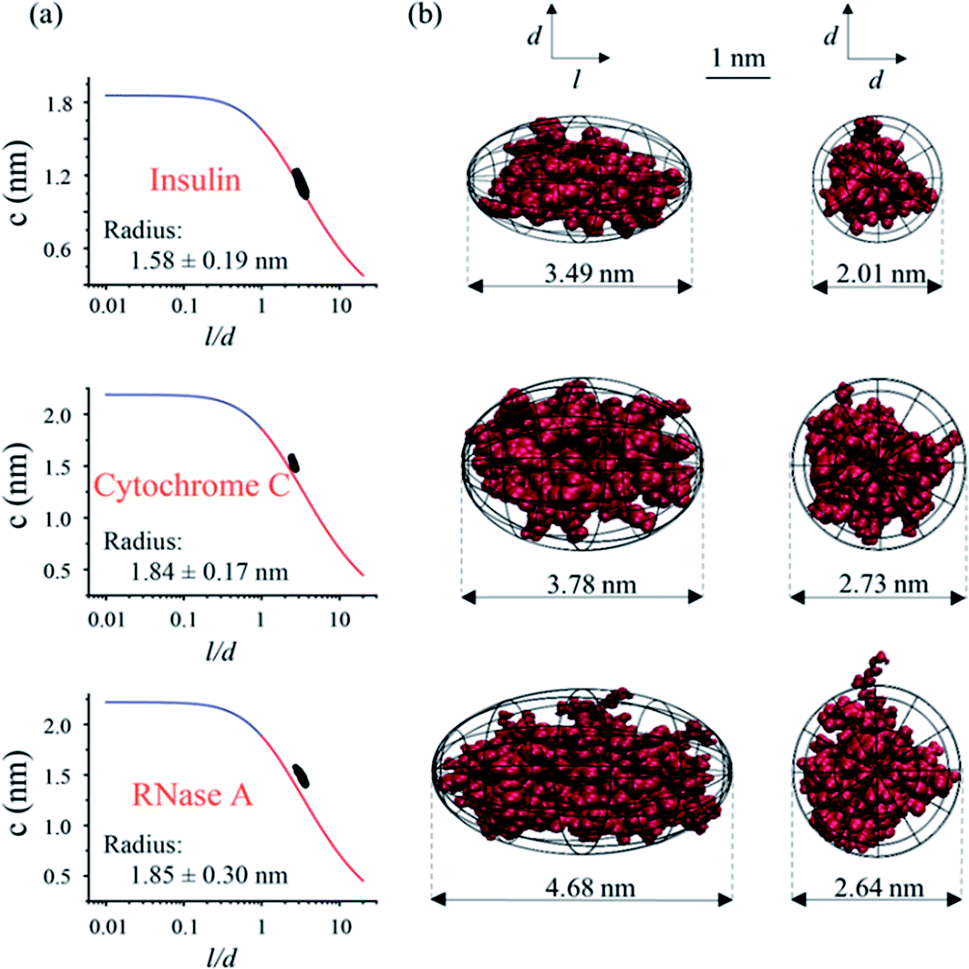
Structure and effective charge characterization of proteins by a mobility capillary electrophoresis based method - Chemical Science (RSC Publishing) DOI:10.1039/C9SC02039J

An Efficient Method for Estimating the Hydrodynamic Radius of Disordered Protein Conformations. - Abstract - Europe PMC

Computing, analyzing and comparing the radius of gyration and hydrodynamic radius in conformational ensembles of intrinsically disordered proteins | bioRxiv
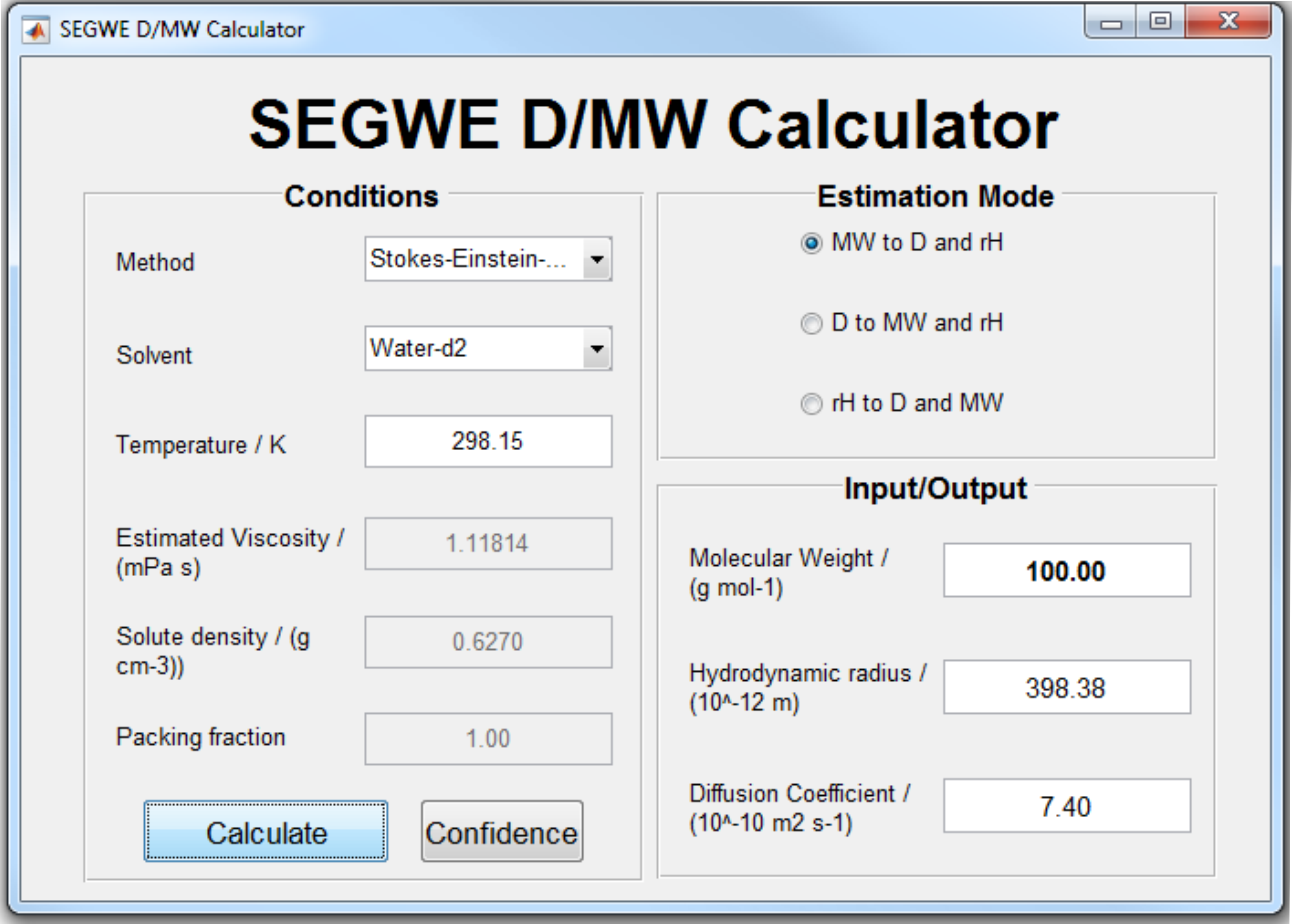
SEGWE calculator: Diffusion coefficient and molecular weight estimation | Manchester NMR Methodology Group
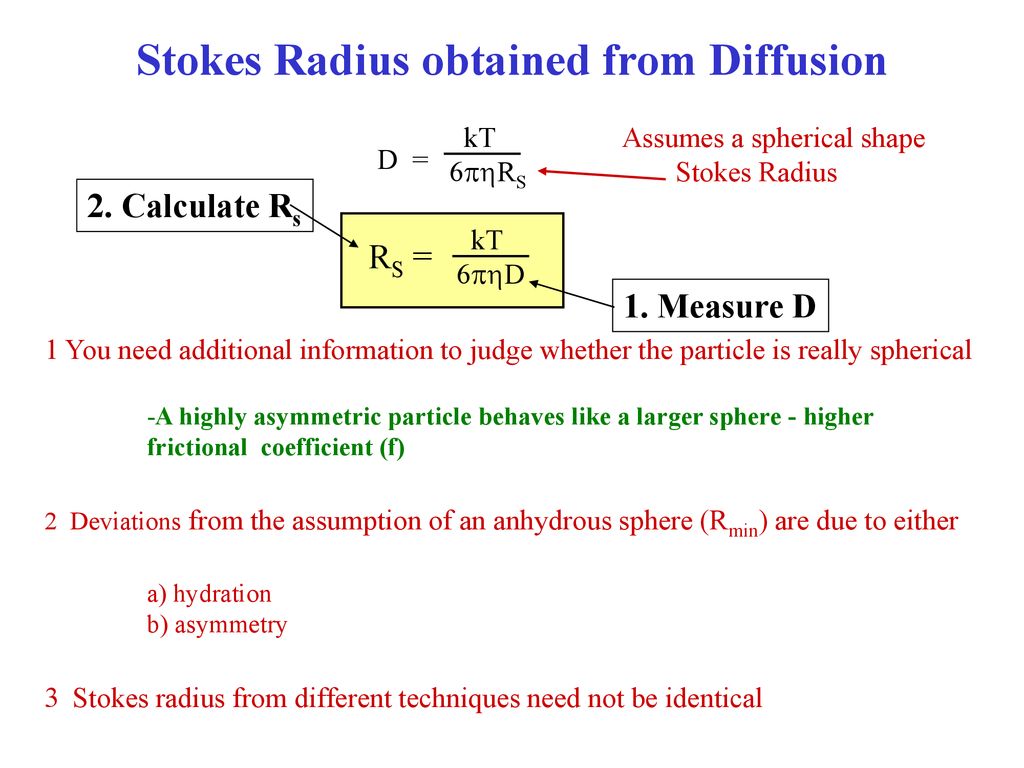

Translational Diffusion: measuring the frictional force on the movement of a macromolecule in solution. A particle under the influence of a constant applied. - ppt download
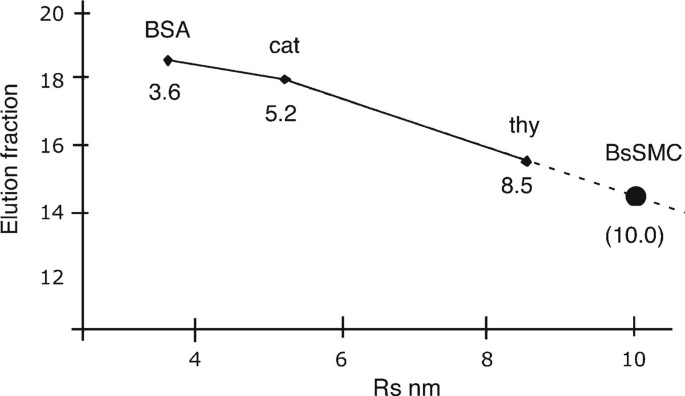
Size and Shape of Protein Molecules at the Nanometer Level Determined by Sedimentation, Gel Filtration, and Electron Microscopy | Biological Procedures Online | Full Text


.jpg)






